A mathematical model of vasoreactivity in rat mesenteric arterioles: I. Myoendothelial communication
Description
To study the effect of myoendothelial communication on vascular reactivity, we integrated detailed mathematical models of Ca2+ dynamics and membrane electrophysiology in arteriolar smooth muscle (SMC) and endothelial (EC) cells. Cells are coupled through the exchange of Ca2+, Cl-, K+, and Na+ ions, inositol 1,4,5-triphosphate (IP3), and the paracrine diffusion of nitric oxide (NO). EC stimulation reduces intracellular Ca2+ ([Ca2+]i) in the SMC by transmitting a hyperpolarizing current carried primarily by K+. The NO-independent endothelium-derived hyperpolarization was abolished in a synergistic-like manner by inhibition of EC SKCa and IKCa channels. During NE stimulation, IP3 diffusing from the SMC induces EC Ca2+ release, which, in turn, moderates SMC depolarization and [Ca2+]i elevation. On the contrary, SMC [Ca2+]i was not affected by EC-derived IP3. Myoendothelial Ca2+ fluxes had no effect in either cell. The EC exerts a stabilizing effect on calcium-induced calcium release-dependent SMC Ca2+ oscillations by increasing the norepinephrine concentration window for oscillations. We conclude that a model based on independent data for subcellular components can capture major features of the integrated vessel behavior. This study provides a tissue-specific approach for analyzing complex signaling mechanisms in the vasculature. Microcirculation (2009) iFirst, 1-25. doi:10.1080/10739680903177539 ADAM KAPELA 1, ANASTASIOS BEZERIANOS 2, AND NIKOLAOS M. TSOUKIAS 1 1-Department of Biomedical Engineering, Florida International University, Miami, Florida, USA; 2-University of Patras, Department of Medical Physics, Patras, Greece
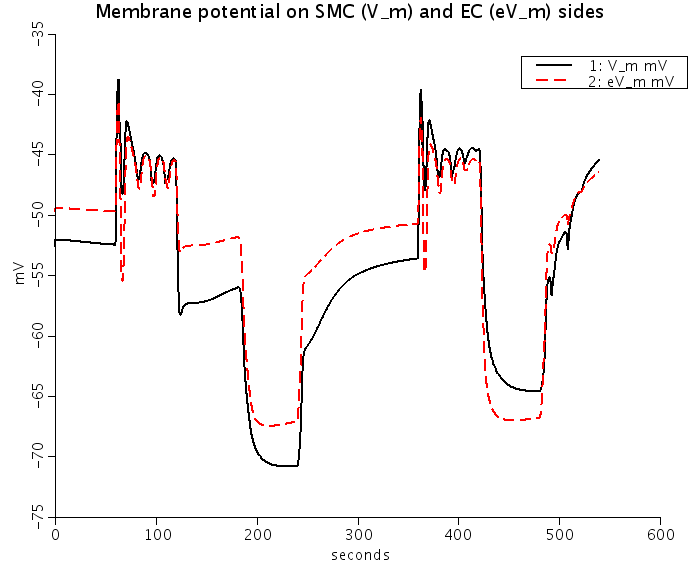
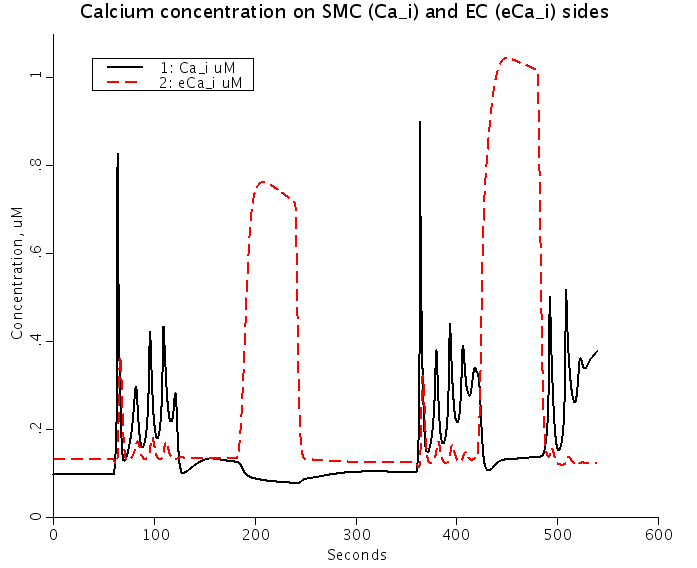
Figure: Predicted Nnorepinephrine-induced and ACh-induced changes of membrane potentials (Top) and intracellular Ca2+ (Bottom) in smooth muscle cells (SMC) (solid lines) and EC (dashed red lines) coupled by the ionic and IP3 fluxes. NE stimulation depolarizes SMC and EC, and increases SMC and EC [Ca2+]i. ACh stimulation repolarizes EC and SMC, increases EC [Ca2+]i, and reduces SMC [Ca2+]i.
Equations
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
We welcome comments and feedback for this model. Please use the button below to send comments:
Kapela A, Bezerianos A, Tsoukias NM: A mathematical model of vasoreactivity in rat mesenteric arterioles: I. Myoendothelial communication, MICROCIRC 16:8,(694-U69), 2009 Kapela A, Bezerianos A, Tsoukias NM.: A mathematical model of Ca2+ dynamics in rat mesenteric smooth muscle cell: agonist and NO stimulation, J Theor Biol 253:238-260, 2008 Silva HS, Kapela A, Tsoukias NM: A mathematical model of plasma membrane electrophysiology and calcium dynamics in vascular endothelial cells. Am J Physiol Cell Physiol 293:C277-C293, 2007
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

