Circulating, multicompartment O2-CO uptake and Hb/Mb binding model based on Erupaka et al. 2010. All table and eq. references are to 2010 paper unless noted. Figures and data referenced here are from Erupaka et al. 2010 paper.
Description
The model written here is described in the Erupaka et al. 2010 paper and is an expansion of a previous model (Bruce et al 2008). A significant enhancement to the 2008 model is the addition of the cardiac compartment. The recirculating model consists of a lung compartment, to handle breathing air into alveolar region, lung capillary compartment, to handle exchange between lung and blood, artery region, handle blood transport to muscles, Skeletal and cardiac muscle, to model O2 consumption, and mixed venous region, to handle blood transport back to lungs. CO and O2 mass balance equations were written for all compartments. CO and O2 entering through the lungs are transported via the major arteries of the arterial blood compartment to the vascular compartments of skeletal muscle (bm1, bm2, bm3), cardiac muscle (bc1, bc2, bc3), and non-muscle tissue (bot). CO and O2 then diffuse into the tissue compartments (m1, m2, c1, c2, ot), driven by pressure gradients. Determining the partial pressures of O2 and CO, and O2 flux in each blood compartment of the model is done by solving the oxyhemoglobin dissociation curve, Haldane’s equation, and blood-to-tissue O2 flux equations, at each time step using an iterative approach to calaculate state variables. The Haldane equation describes the competition of CO and O2 for hemoglobin (Hb) and Mb binding sites.
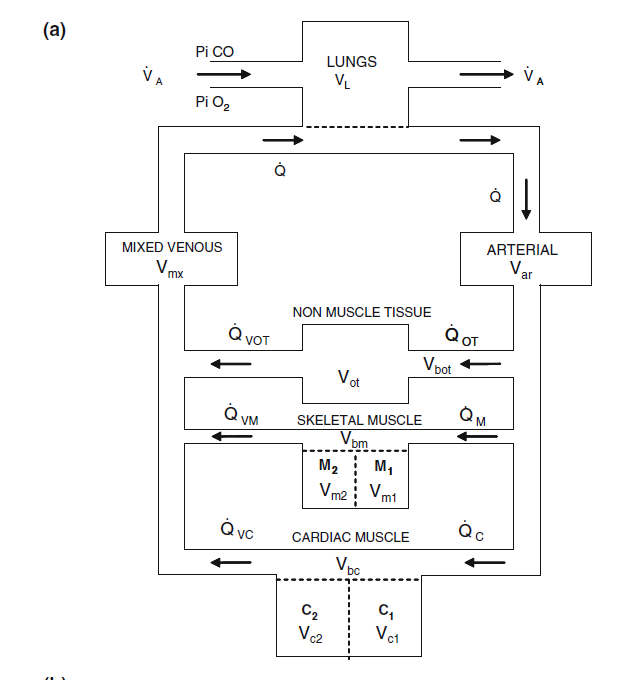
Figure 1: Diagram of model, showing basic flow of blood/gases and compartments. Taken from Figure 1a of Erupaka et al. 2010 paper listed below. Please see paper or model code for more details.
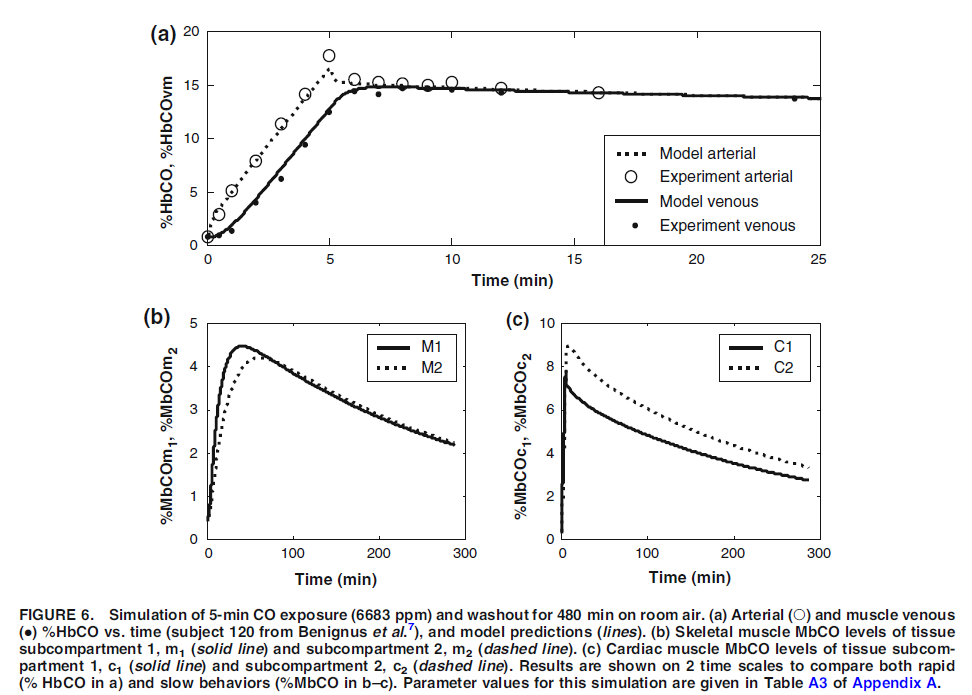
Figure 2 (above): Figure taken directly from figure 6 of Erupaka et al. 2010 paper.
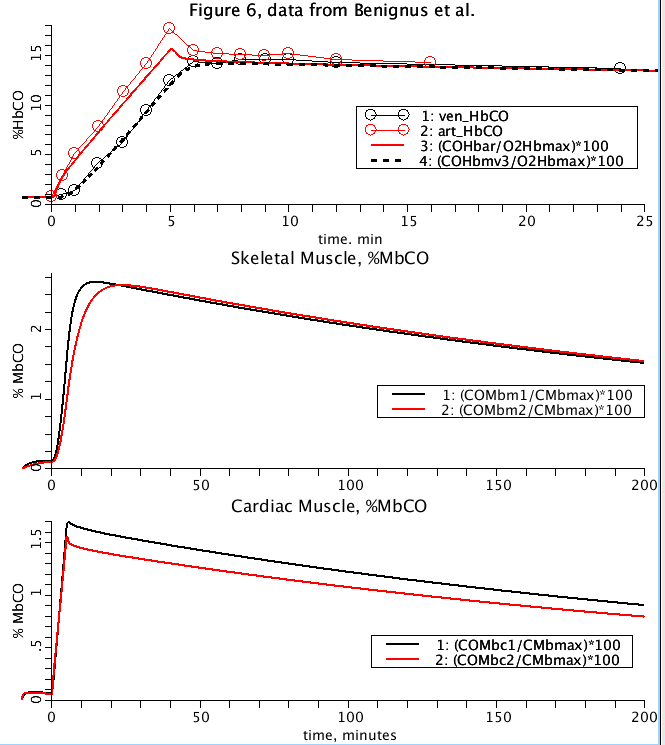
Figure 3: JSim model simulation attempting to reproduce figure 6 from Erupaka et al. 2010 and fit data from Benignus et al. 1994.
JSim model implimentation comments
- Currently unable to reproduce difference in bound %HbCO between arterial blood and muscle venous blood See Fig 6 parameter set and plotpage '2010_fig6'.
- Our model behaviour in the Cardiac muscle tissue regarding %MbCO levels is not as pronounced beteen compartment 1 and 2 and Comp 1 and 2 percentages 'cross over'. See fig 6 and fig 7 parameter sets and plots.
- Replaced ep and ec regions of Bruce et al. 2010's model for a compartment called Lcap (Lung capillary compartment) Want a better model of diffusion of gasses from lung to aveolar capillaries. Removed circulatory transport delays. Use ODEs to describe transport of gasses into and out of the lung.
- Added a term to the alveolar gas equations for CAO2 to take into account CO2 (avg term). Use of this maintains a PAO2 of around 100 Torr rather than a partial pressure near PIO2. May need a more detailed aveolar gas equation to take into account large changes in FIO2 and the diluting effect of N2. Going forward we will want to add CO2 equations. Ref: Hlastala and Berger, Physiology of Respiration, 2001, p65-80.
- Model takes approximately 10 min to reach steady-state, Do not make any changes from initial conditions until approx 10 min has elapsed (param sets usually have t.min = -10 min).
- JSim has trouble finding solutions to the implicit equations and changing some parameters by more than 20% may causes 'zero-finder' errors and simulation will be unable to proceed. The model is not robust.
Equations
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
We welcome comments and feedback for this model. Please use the button below to send comments:
- KINNERA ERUPAKA, EUGENE N. BRUCE, and MARGARET C. BRUCE, Prediction of Extravascular Burden of Carbon Monoxide (CO) in the Human Heart , Annals of Biomed Eng, Vol. 38(2), Feb 2010, pp. 403–438
- Bruce, EN, Bruce MC, and Erupaka K. Prediction of the rate of uptake of carbon monoxide from blood by extravascular tissues . Respir. Physiol. Neurobiol. 161(2):142–159, 2008
- Bruce, E. N., and M. C. Bruce. A multicompartment model of carboxyhemoglobin and carboxymyoglobin responses to inhalation of carbon monoxide. J. Appl. Physiol. 95(3):1235–1247, 2003.
- Benignus, V. A., M. J. Hazucha, M. V. Smith, and P. A. Bromberg. Prediction of carboxyhemoglobin formation due to transient exposure to carbon monoxide. J. Appl. Physiol. 76(4):1739–1745, 1994.
- Burge, C. M., and S. L. Skinner. Determination of hemoglobin mass and blood volume with CO: evaluation and application of a method. J. Appl. Physiol. 79(2):623–631, 1995.
- Hlastala MP, Berger AJ, Physiology of Respiration, Oxford Univ Press, NY,NY, 2001, chap 4(65-80)
- Model #153, Highly-integrated human model
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

