Aspirin and Salicylate circulate in the plasma and are converted to salicylurate in liver mitochondria prior to clearance, mainly as salicylurate.
Description
Diverse data sets can yield reliable information through
mechanistic modeling: salicylic acid clearance
by
Raymond GM and Bassingthwaighte JB.
NSR Model 369 provides 3 models each fitting of data from the
same three sources as here:
1. Exponential Washout or Half-life for clearance
2. Michaelis-Menten reaction in plasma.
3. Enzyme kinetics, fully reversible reactions for substrate and product
This MODEL 377 CONSIDERS THE REACTION TO OCCUR IN MITOCHONDRIA (LIVER)
BUT CLEARNCE TO BE FROM THE BLOOD. The mito volume is set to 1/1000 of the
blood volume, so compared to the Model 369 solutions the kinetic parameters
are the same, except that a large Etot and therefore Vmax in the mito
is used to give the same total reaction capacity.
kon1 koff2
S + E ------> SE ------> E + P
<------ <------
koff1 kon2
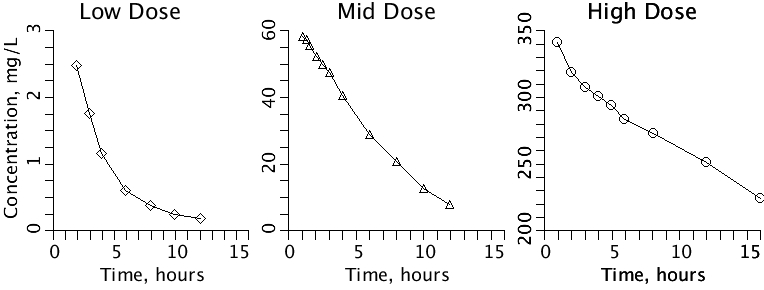
Figure 1: Fig01_DataPlot:
The three sets of digitized data from the references
are in DataSets lowdose, middose, and high dose. These files
will be referenced in the plots and used for optimizing
parameters.
Figure 2: EnzSatur:
Plotted versus time are the blood and intramitochondrial concns
and the fractional saturation of the enzyme inside the mito. The blood data
are shown by the symbols.
Figure 3: FitEnzMito:
For each dose level the plasma concn (red) and the intramito concn (blue)
are plotted against the blood data for the three data sets. The concn of
product is green: solid lines for inside the mito and dashed lines for blood.
Product blood levels are lower because the only clearance is from the blood.
Salicylate blood levels are always higher than mito levels since direct
salicylate clearance from blood is low, and inside the mito it is being
converted to product salicylurate. Parameters as in Table 3, right 3 columns
of Raymond and Bassingthwaighte Brit J Pharm Res 2015.
Equations
None.
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
We welcome comments and feedback for this model. Please use the button below to send comments:
Aarons L, Hopkins K, Rowland M, Brossel S, Thiercelin JF. Route of administration and sex differences in the pharmacokinetics of aspirin, administered as its lysine salt. Pharmaceutical Res. 1989; 6: 660-666. Benedek IH, Joshi AS, Pieniaszek HJ, SY King SY and Kornhauser DM. Variability in the pharmacokinetics and pharmacodynamics of low dose aspirin in healthy male volunteers. J. Clin. Pharmacol. 1995; 35: 1181-1186. Prescott LF, Balali-Mood M, Critchley JA, Johnstone AF, Proudfoot AT. Diuresis or urinary alkalinisation for salicylate poisoning? Br Med J (Clin Res Ed). 1982; November 13; 285(6352): 1383–1386.
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

