This model is a place-holder for the three separate models, HalfLife, Enzyme, and BriggsHaldane. It is an example of reproducible models leading to reproducible science. Run the SalicylicAcidClearance model first.
Description
This is the project file for all the figures and tables in Raymond, GM and Bassingthwaighte JB. Reproducible salicylic acid clearance models. Three data sets for acid clearance have been combined covering low, medium, and high dose ranges of aspirin. The averaged plasma concentrations are model by three different models, (1) HalfLife, (2) enzyme, and (3) Briggs-Haldane Michaelis-Menten. Various pitfalls leading to the non-reproducibility of the models are explored. Paraphrasing Box (1976)“All models are wrong but some are useful,” it is concluded, "If it is true that 'All models are wrong,' then models that are not reproducible are not even useful. Use the directions under the README tab in conjunction with the paper to reproduce all figures and tables in the publication.
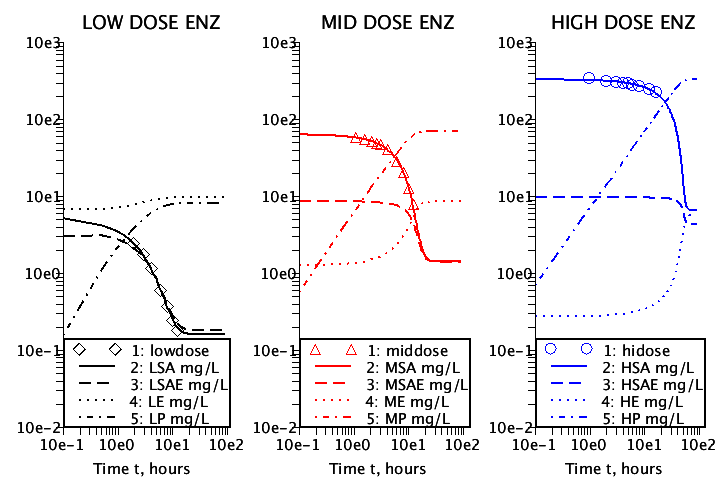
Figure 7. The complete enzyme solutions. The symbols are the data; the solid lines are the fits to the data; the dashed lines are the concentrations of the SA-enzyme complex; the dotted lines are the concentrations of the free enzyme; and the dashed- dotted lines are the metabolized product. The low dose concentrations illustrate Michaelis-Menten kinetics where the substrate and the substrate-enzyme complex are in equilibrium because of under saturation of the available enzyme concentration. The mid and high dose concentrations illustrate Briggs-Haldane kinetics where the rate of change of the substrate-enzyme complex is very small because of over saturation of the available enzyme concentration.
Equations
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
We welcome comments and feedback for this model. Please use the button below to send comments:
Aarons L, Hopkins K, Rowland M, Brossel S, and Thiercelin JF. Route of administration and sex differences in the pharmacokinetics of aspirin, administered as its lysine salt. Pharmaceutical Res 6: 660-666, 1989. Bassingthwaighte JB, Butterworth E, Jardine B, and Raymond G. Compartmental modeling in the analysis of biological systems. In "Computational Toxicology: Volume 1", edited by Brad Reisfeld and Arthur N Mayeno. Methods in Molecular Biology 929: 391-438, 2012. Benedek IH, Joshi AS, Pieniaszek HJ, SY King SY and Kornhauser DM. Variability in the pharmacokinetics and pharmacodynamics of low dose aspirin in healthy male volunteers. J. Clin. Pharmacol 35: 1181-1186, 1995. Box GEP. Science and statistics. J. Am. Stat. Assoc. (1976) 71:791–799. Chan IS, Goldstein AA, and Bassingthwaighte JB. SENSOP: a derivative-free solver for nonlinear least squares with sensitivity scale. ABME, (1993) 22:621-631. Hutt AJ, Caldwell L, and Smith RL. The metabolism of aspirin in man: a population study. Xenobiotica, 1986, 16:239-249. Kuehl GE, Bigler J, Potter JD, Lampe JW. Glucuronidation of the aspirin metabolite salicylic acid by expressed UDP-glucuronosyltransferases and human liver microsomes. Drug Metab Dispo. 2006 Feb;34(2):199-202. Epub 2005 Oct 28. Prescott LF, Balali-Mood M, Critchley JA, Johnstone AF, Proudfoot AT. Diuresis or urinary alkalinisation for salicylate poisoning? Br Med J (Clin Res Ed). 1982 November 13; 285(6352): 1383–1386.
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

