The Sangren-Sheppard model (Bull Math Biophys 15: 387-394, 1953) for the exchange of a labeled substance between a liquid flowing in a vessel and an external compartment. Similar to BTEX20 with no diffusion
Description
The Sangren-Sheppard model may be seen as a varient of the btex20_pde
model. Instead of specifying Vp, Visfp, and PSg, three alternative
parameters are specified, (1) tcap, the capillary transit time, (2) gamma,
the ratio of Visfp to Vp, and (3) psf, the ratio of of PSg to Fp (flow).
Vp, Visfp, and PSg are then given by
Vp=Fp*tcap, Visfp=gamma*Vp, and PSg=psf*Fp. Once these parameters
are calculated, the model is the same as btex20_pde with out diffusion.
This is a model of a "tissue cylinder" consisting of a capillary
plasma region and an interstitial fluid (isf) region. The model is
multi-segmented in a single spatial dimension to solve the convection-
diffusion equation with exchange and consumption. It is modeled as a
differential equation coupled with an ordinary differential equation.
The convecting plasma region has volume Vp, flow Fp, first order
consumption coefficient Gp. The interstitial fluid (isf) region has volume
Visfp, first order consumption coefficient Gisf. The name of the
isf volume, Visfp, is derived from Visf' where the prime is rendered as p.
The isf volume is a volume of distribution which might or might not
correspond to a physical volume. It is referred to as a virtual volume.
PS is the permeability surface product of the membrane. It is also
known as PScap (PS for the CAPillary region) and PSg (the PS for the
Gap between endothelial cells which allows for the passive exchange of
material between the plasma and the isf. In this model, the PS is denoted
PSg. In other nomenclature it is denoted PSc where c stands for "cleft",
a synonym for gap.
Units are per gram of tissue (physiological) so that a single sanshe
unit can model a homogeneously perfused organ.
Instantaneous mixing is assumed in each individual segment in each
region. Passive exchange between plasma and isf regions has conductance
PSg.
This model can be adapted and expanded to represent multi-capillary
models in serial and parallel configurations.
Equations
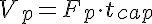
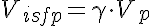
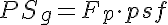
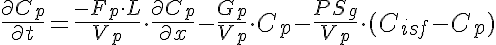
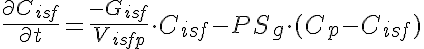
Initial Conditions:
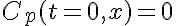
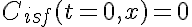
Boundary Conditions:
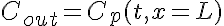
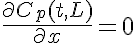
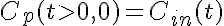
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
- No SBML translation currently available.
- Information on SBML conversion in JSim
We welcome comments and feedback for this model. Please use the button below to send comments:
Sangren WC and Sheppard CW. A mathematical derivation of the exchange of a labeled substance between a liquid flowing in a vessel and an external compartment. Bull Math Biophys 15: 387-394, 1953 (This gives an analytic solution for the two-region model.) Goresky CA, Ziegler WH, and Bach GG. Capillary exchange modeling: Barrier-limited and flow-limited distribution. Circ Res 27: 739-764, 1970. (This gives another derivation of the analytical form, and uses the model in both single and multicapillary models. Bassingthwaighte JB. A concurrent flow model for extraction during transcapillary passage. Circ Res 35: 483-503, 1974. (This gives numerical solutions, which are faster than the analytic solutions, and imbeds the model in an organ with tissue volums conserved, and with arteries and veins.) Guller B, Yipintsoi T, Orvis AL, and Bassingthwaighte JB. Myocardial sodium extraction at varied coronary flows in the dog: Estimation of capillary permeability by residue and outflow detection. Circ Res 37: 359-378, 1975. (Application to sodium exchange in the heart.) Poulain CA, Finlayson BA, Bassingthwaighte JB. Efficient numerical methods for nonlinear-facilitated transport and exchange in a blood-tissue exchange unit, Ann Biomed Eng. 1997 May-Jun;25(3):547-64.
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

