Comparison of 1-sided and 2 sided Michaelis-Menten transporters in a two compartment model with flow.
Description
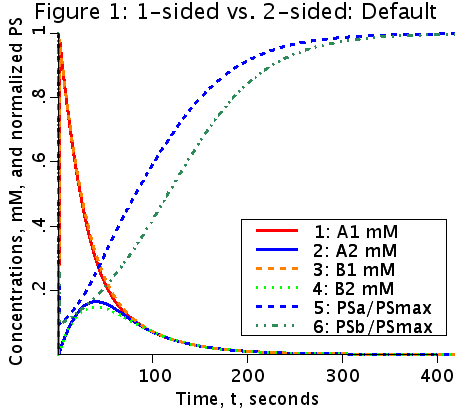
Figure 1: Concentrations of A1, A2, B1, and B2 are plotted as functions of time. The normalized values of PS (PSa/PSmax and PSb/PSmax) are also plotted as functions of time.
Two types of a saturable Michaelis-Menten transporter are considered in this two compartment model with flow--a one sided transporter for solute A and a two-sided transporter for solute B. The model for solute A is cis- side driven. Concentration of A in V1, A1, determines the fractional saturation, PSa/PSmax, where PSmax is Vmax/Km, and PSa = PSmax/(1 + A1/Km). The model for solute B is cis-trans driven. Concentration of B in both V1 and V2, B1 and B2 respectively, determine the fractional saturation, where PSmax is Vmax/Km and PSb = PSmax/(1 + B1/Km + B2/Km).
Equations
One sided Michaelis-Menten Transporter
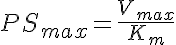

Ordinary Differential Equations
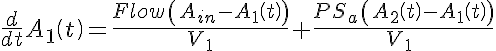
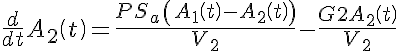
Initial Conditions
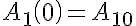
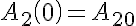
Two sided Michaelis-Menten Transporter
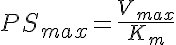
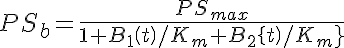
Ordinary Differential Equations
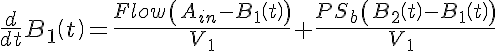
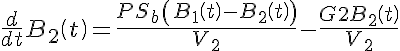
Initial Conditions
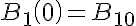
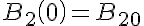
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
We welcome comments and feedback for this model. Please use the button below to send comments:
Klingenberg M. Membrane protein oligomeric structure and transport function. Nature 290: 449-454, 1981. Stein WD. The Movement of Molecules across Cell Membranes. New York: Academic Press, 1967. Stein WD. Transport and Diffusion across Cell Membranes. Orlando, Florida: Academic Press Inc., 1986. Wilbrandt W and Rosenberg T. The concept of carrier transport and its corollaries in pharmacology. Pharmacol Rev 13: 109-183, 1961. Schwartz LM, Bukowski TR, Ploger JD, and Bassingthwaighte JB. Endothelial adenosin transporter characterization in perfused guinea pig hearts. Am J Physiol Heart Circ Physiol 279: H1502-H1511, 2000.
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

