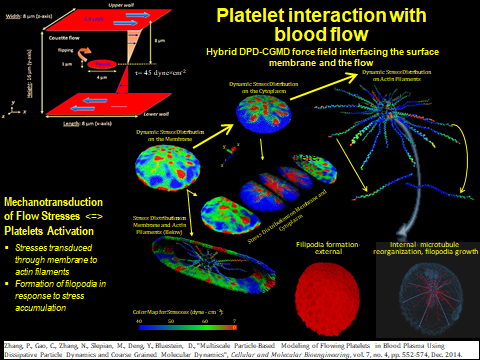
We will use an integrated Dissipative Particle Dynamics (DPD) and Coarse Grained Molecular Dynamics (CGMD) approach that allows platelets to continuously change their shape and synergistically activate by a biomechanical transductive linkage chain, interact with other blood constituents and clotting factors, aggregate, and interact and adhere to the blood vessels and devices. In this multiscale model, a mechanotransduction CGMD bottom platelet activation model is embedded into a DPD blood flow top model. The dynamic stresses of the macroscale model are interactively translated to the micro- to nanoscale model of the intra-platelet associated intracellular events. The model predictions are validated in vitro in a carefully designed set of experiments.
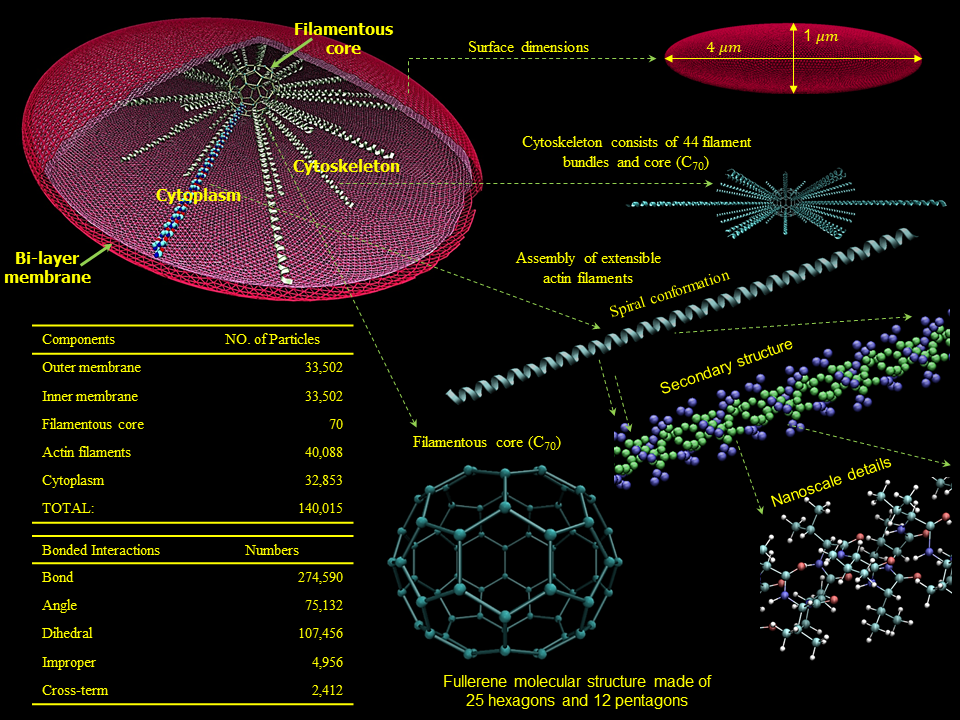
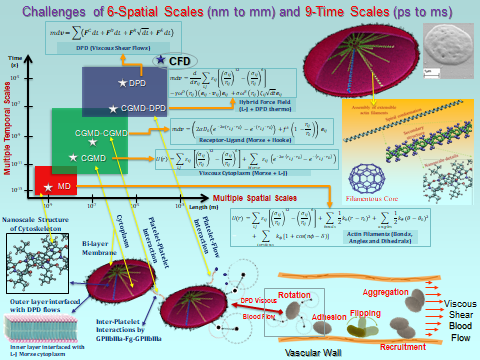
We have developed a mechanotransduction model of platelet mediated-thrombosis, where a top/macro-scale model of flow-induced thrombogenicity using DPD at the µm-length and ms-time scales, in which multiple flowing platelets interact with each other and blood vessel walls or devices, and is fully coupled with a bottom/micro-scale model using CGMD at the nm-length and ps-time scales, in which platelets with multiple intracellular constituents evolve during activation as platelet lose their quiescent discoid shape and filopodia grow. The top and bottom models are interfaced such that the hemodynamics interactively respond to platelet shape change upon activation and platelet aggregation, and allow a thrombus to form. The model includes intraplatelet constituents (filamentous actin and microtubular cytoskeleton, cytoplasm), bilayer plasma membrane, and GPIIb-IIIa and GPIbα receptors for interaction with fibrinogen and von Willebrand factor during aggregation and adhesion, respectively. The dynamics and morphologies of the platelet model have been carefully validated in vitro for their mechanical properties, and flow-mediated platelet activation and aggregation. The model is currently undergoing development and in vitro validation for adhesion. This model can also be used to evaluate the effect of modulating platelet mechanical properties via antiplatelet agents. These data will be used to fine tune the large number of model parameters involved in this multiscale simulation and for validating the model predictions.
All model aspects are validated in vitro in a series of carefully designed experiments characterizing the mechanical properties of platelets and using blood flow experiments where conditions leading to flow induced platelet activation will be replicated, as well as experiments where platelet-wall and platelet device interactions will be measured and where platelets will be pretreated with modulating agents. We use platelets obtained from consenting healthy adult volunteers under Stony Brook University and University of Arizona IRB-approved protocols. Validation involves tools such as dielectrophoresis-mediated electrodeformation to measure mechanical properties, cone-plate-Couette viscometry and microfluidics to induce platelet activation, aggregation, and adhesion, scanning electron microscopy (SEM) to evaluate morphological details of activated platelets, and high-resolution and high-framerate DIC microscopy to capture platelet morphological changes and dynamics under shear flow.
Gupta, P.; Zhang, P.; Sheriff, J.; Bluestein, D.; and Deng, Y. A Multiscale Model for Recruitment Aggregation of Platelets by Correlating with In Vitro Results. Cellular and Molecular Bioengineering, 12: 327-343. 2019.
Zhang, P.; Zhang, L.; Slepian, M. J.; Deng, Y.; and Bluestein, D. A Multiscale Biomechanical Model of Platelets: Correlating with In-Vitro Results. Journal of Biomechanics, 50: 26–33. 2017.
Gao, C.; Zhang, P.; Marom, G.; Deng, Y.; and Bluestein, D., Reducing the Effects of Compressibility in DPD-based Blood Flow Simulations through Severe Stenotic Microchannel, Journal of Computational Physics, vol. 335, pp. 812-827, 2017.
Zhang, P.; Zhang, N.; Gao, C.; Zhang, L.; Gao, Y.; Deng, Y.; and Bluestein, D., Scalability Test of Multiscale Fluid-Platelet Model for Three Top Supercomputers, Computer Physics Communications, vol. 204, pp. 132-140, 2016.
Zhang, P.; Zhang, N.; Deng, Y.; and Bluestein, D., A Multiple Time Stepping Algorithm for Efficient Multiscale Modeling of Platelets Flowing in Blood Plasma, Journal of Computational Physics, vol. 284, pp. 668-686, 2015.
Pothapragada, S.; Zhang, P.; Sheriff, J.; Livelli, M.; Slepian, M. J.; Deng, Y.; and Bluestein, D. A Phenomenological Particle-Based Platelet Model for Simulating Filopodia Formation During Early Activation. International Journal for Numerical Methods in Biomedical Engineering, vol. 31, no. 3, pp. 1-16. 2015.
Zhang, P.; Gao, C.; Zhang, N.; Slepian, M. J.; Deng, Y.; and Bluestein, D. Multiscale Particle-Based Modeling of Flowing Platelets in Blood Plasma Using Dissipative Particle Dynamics and Coarse Grained Molecular Dynamics. Cellular and Molecular Bioengineering, vol. 7, no. 4, pp. 552-574. 2014.
Bluestein, D., Soares, J.S., Zhang, P., Gao, C., Pothapragada, S., Zhang, N., Slepian, M.J., Deng, Y., Multiscale Modeling of Flow Induced Thrombogenicity With Dissipative Particle Dynamics and Molecular Dynamics, Journal of Medical Devices, vol. 7, issue 4, pp. 024502-024503, 2014.
Zhang, N., Zhang, P., Kang, W., Bluestein, D., Deng, Y., Parameterizing the Morse potential for coarse-grained modeling of blood plasma, Journal of Computational Physics, vol. 257, Part A, pp. 726-736, 2014.

