Michaelis-Menten Reaction Sequence for solutes A to E in a stirred tank, with lagged input into flowing reactor with constant volume, Vol, and step jumps in flow and in reaction rates. This is the third in a series of four models to account for substrate capacitance in enzymatic networks. The first model is Linear.Reaction.Sequence (Model #0423).
Description
A reaction sequence A-->B-->C-->D-->E can be represented many ways to approximate the biological form of the reactions.
This version is the next to simplest: Michaelis-Menten unidirectional reaction clearances, giving progress curves
as in a bioreactor, but with additional factor of a flow through the mixing chamber. The flow term is first order
for all solutes in the sequence, but the reaction terms are saturable MM, Vmax*A/(Km+A).
The inflowing initial solute A, the flow, ml/sec, Flow*CinA, equals the sum of the outflow clearance plus the
last reaction E--> ?, i.e. G*E, then the outflow concentration of A in the steady state goes to 50% of CinA
(Do this by setting Flow1 = Flow2 = 0.05 ml/sec and setting GA1 and GA2 also to 0.05 and run the program. The
other reaction rates, to form C, D, and E are set identidal to that to form B from A, the result is that steady state
concentrations for A, B, etc go to 0.5, 0.25, 0.125, 0.0625, and 0.03125, all at one half of its predecessor.
This program illustrates the transient delays between steady state, and the form of the transients.
With MM kinetics there is really no enzyme there, and no binding of the reactants in the process of the reaction.
All of the delay is due to the combination of flow and the reaction, exactly similar to the first order reaction
kinetic model (Model #432). The initial conditions are zero for all solutes
so the time constant for the initial entry would be simply the volume divided by flow, Vol/Flow1, in seconds,
if there were no reaction. The reaction, augmenting the disappearance of A, shortens the time constant so that
it is Vol/(GA + Flow1).
The second transient is due to a step increase (or decrease) in flow at time TFjump (at t=30 s with this
parameters set (LagMM.181026). The third transient is a step change in the reaction rates at time TGjump.
The Verification Process is the same as for the linear kinetics, and which does verify the code whn the solute
concnetrations are 1 mM hen the Km was
The solution to the differential equation for solute A is accomplished using the numerical solvers,
but it is also solvable analytically. The three transients, at t = 0, t= 30 s and t=60 s, are expressed in three
analytical equations. Their sum fits exactly the numerical solutions to the systems equations after the effects of
lag TauA on the conentration of A in the compartment. VERIFICATION!
Try this out using the difference ODE solvers: it turns out the DOPRIS5 (an advanced RungeKutta algorithm)
gives a more precise fit to the analytical solution than either of the more powerful stiff solvers, CVODE or
Radau. At the steep parts of the transients DOPRIS5 is still good to 7 decimal digits, while the other supposedly
superior solvers, CVODE and Radau are good only 5 or 6 decimal digits. In the steady state they all give the same
correct answers.
For solutes B through E, the analytical solutions are more complex, and have not been developed. For these
it is much faster to use the numerical solutions to the ODEs; the analytical solutions for solute E would be,
because of their complexity, much slower to compute, and maybe not even as accurate as the numerical solutions,
even for this rather simple model system. What we know is that the steady state solution match exactly to the
predicted steady state values for A through E.
This model is a variant on Michaelis-Menten that approximates the effect of solute binding to the
enzymes (Model #425). That is done to correct the inadequate kinetics of M-M expressions. This family of models
is designed for use as a set of tools to determine the magnitude of the buffering capacitance and consequent
delay in transient responses in enzyme systems. In this case RUN LOOPS to double Etot and halve k2A to keep
the flux the same.
The basis for this series of models to account for substrate capacitance in enzymatic networks
is first the model Linear.Reaction.Sequence (Model #423); the second Model (#424) is MM.Reaction.Sequence,
and this is the third, MM.Lag.Reactio.Sequence (Model #425). The fourth, (Model #426)
adds the on- and off kinetics for a single uncomplicated reaction, Enzyme.React.Seq.Full.Kinetics.
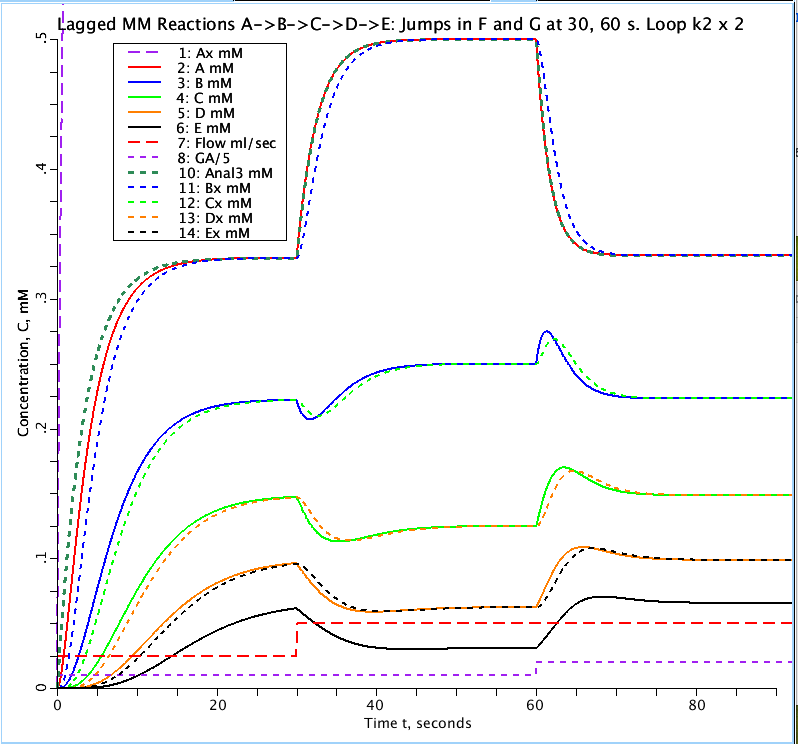
Figure: Progress curves for a sequence of reactions from substrate A to E in a compartment with flow through the compartment. All substrates have initial concentration of zero with A in (CinA) set to 1 mM. The lag terms Ax, Bx, Cx, Dx, Ex (mM) are the added terms use to approximate a delay in free substrate due to enzyme binding. Flow doubles from 0.025 to 0.05 ml/sec at t= 30 sec. Consumption (the flow of fluid cleared of substrate per unit time) doubles from 0.05 to 0.1 ml/sec at t= 60 sec. GA is the consumption of substrate A and Anal3 is the analytical solution for substrate concentration A as a function of time and matches A(t) after the lag is taken into account.
Equations
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
- No SBML translation currently available.
- Information on SBML conversion in JSim
We welcome comments and feedback for this model. Please use the button below to send comments:
Easterby, JS. A generalized theory of the transition time for sequential enzyme reactions.
Biochem J. 199: 155-161, 1981.
Cascante M, Melendez-Hevia E, Kholodenko B, Sicilia J, and Kacser H. Control analyis
of transit time for free and enzyme-bound metabolites: physiological and
evolutionary significance of metabolic response times. Biochem J 308: 895-899, 1995.
Bassingthwaighte JB. Capacitance in metabolic netowrks. 2019 (in prep for submission to Biophys J)
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

