Flow with axial dispersion through a one-region pipe of uniform cross-section. Statistics on inflow and outflow concentration curves */
Further reading: Distributed Blood Tissue Exchange Models Explained
Description
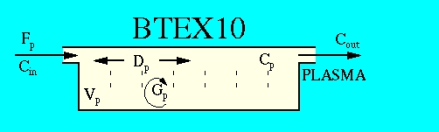
The partial differential equation models flow into, through and out of a pipe with plug flow and axial dispersion (diffusion) along the x-axis and instantaneous radial dispersion so that concentration is uniform across the cross-section at each x-position. Consumption,Gp, equivalent to loss by a first order reaction or loss by permeation is a uniform fraction per unit time along the pipe. (This can be modified by making G a function of concentration, Gp(Cp) or of position, Gp(x).) Flow is constant, as are all the other parameters.The boundary conditions are (1) At the inflow, the diffusion coefficient, Dp, cm^2/s, times the spatial gradient in concentration, dC/dx, balances the difference between the inflow concentration and the concentration Cp just inside; (2) At the outflow, the gradient dC/dx is set to zero, as if reflecting from an impermeable surface, so that mass is lost into the outflow only by flow, Cout = Cp(x=L,t). LIMITATIONS: This model cannot approximate Newtonian parabolic flow, where the response to a flow-proportiaonal cross-sectional pulse labeling at the inflow would give a sharp upstroke and peak at 1/2 the mean transit time and then, in the absence of axial dispersion, diminish in proportion to 1/t^2. See Gonzalez-Fernandez (1962) on this point.
Equations
Differential Equations
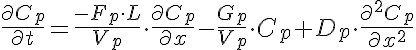
Left Boundary Conditions
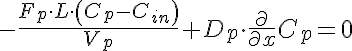
Right Boundary Conditions
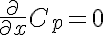 ,
, 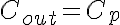 .
.
Initial Conditions
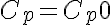 or
or
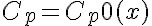
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
- No SBML translation currently available.
- Information on SBML conversion in JSim
We welcome comments and feedback for this model. Please use the button below to send comments:
W.C. Sangren and C.W. Sheppard. A mathematical derivation of the exchange of a labelled substance between a liquid flowing in a vessel and an external compartment. Bull Math BioPhys, 15, 387-394, 1953. Gonzalez-Fernandez JM. Theory of the measurement of the dispersion of an indicator in indicator-dilution studies. Circ Res 10: 409-428, 1962. C.A. Goresky, W.H. Ziegler, and G.G. Bach. Capillary exchange modeling: Barrier-limited and flow-limited distribution. Circ Res 27: 739-764, 1970. J.B. Bassingthwaighte. A concurrent flow model for extraction during transcapillary passage. Circ Res 35:483-503, 1974. B. Guller, T. Yipintsoi, A.L. Orvis, and J.B. Bassingthwaighte. Myocardial sodium extraction at varied coronary flows in the dog: Estimation of capillary permeability by residue and outflow detection. Circ Res 37: 359-378, 1975. C.P. Rose, C.A. Goresky, and G.G. Bach. The capillary and sarcolemmal barriers in the heart--an exploration of labelled water permeability. Circ Res 41: 515, 1977. J.B. Bassingthwaighte, C.Y. Wang, and I.S. Chan. Blood-tissue exchange via transport and transformation by endothelial cells. Circ. Res. 65:997-1020, 1989. Poulain CA, Finlayson BA, Bassingthwaighte JB.,Efficient numerical methods for nonlinear-facilitated transport and exchange in a blood-tissue exchange unit, Ann Biomed Eng. 1997 May-Jun;25(3):547-64.
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

