Barrier-limited model of Goresky, using Finite difference method
Theory
Single capillary
The concentrations in plasma, C1 and in tissue, C2, follow the following system of two differential equations in time, t, and space, x:
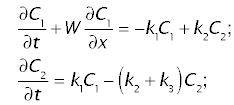
where W is the linear velocity of the tracer the sinusoidal plasma, k1 and k2 are transfer coefficients or rate constants, defining the transfer of tracer between the plasma and tissue, and k3 is the transfer coefficient of sequestration of tracer due to metabolism and/or biliary excretion.
The boundary conditions are
- at t = 0: C1 = 0 and C2 = 0;
- at x = 0: C1 = Cin
where Cin is the tracer concentration at the inflow of the organ. For quasi-instantaneuse injection, administration of tracer will be represented by the Dirac impulse function: Cin(t) = δ(t). In order to make calculations easier, concentrations are normalized by dividing through the injected amount, and distances are normalized by dividing through the linear velocity W, thus assuiming W = 1.
Whole organ
The tracer concentration leaving the whole organ, Cdiff, is obtained as the flow-weighted average of the cocentration in the individual sinusoids. Let f(τc) be the fraction of total organ blood flow emerging from paths with sinusoidal transit times between τc and τc + dτc. The mixed outflow concentration from the whole organ, C(t), will then be the flow-weighted average of the outflow concentrations, according to the integral
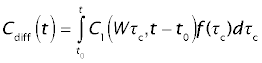
For the reference tracer (sucrose), the normalized concentration at the portal vein is
Cref = f(τc + t0)
Thus,
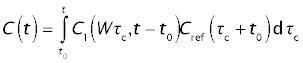
Closed solution
The closed ("analytical") solution for the whole-organ response is
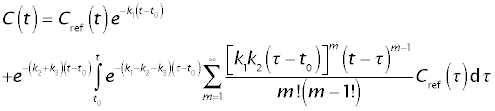
The solution consists of two components:
- The throughput component, e-k1(t - t0) Cref(t), representing tracer that remained in the extracellular space.
- The returning component, repesented by the second line of the above equation, representing tracer that has entered the tissue at least one and has returned to the extracellular space.
Calculations
Parameter sets
There are five parameter sets in this JSim model project file:
- Gal1: Galactose experiment with no galactose infused
- Gal2: Galactose experiment with high galactpse concentration
- Pal1: Palmitate acid experiment (normal control)
- Pal2: Palmitate acid experiment with infusion of α-bromopalmitate
- Rb: Rubidium experiment
To change the parameter set:
- Select Load project parameter set from the ParSet pulldown menu
- Choose the desired parameter set. This will automatically change the paremters as well as the data for the reference curve.
- Click on "Run" to use the new parameter set.
- In order to show the correct data in the plot, select the data set and the tracer from the pulldown menus labeled "data".
Closed solution
Select model "ClosSol" from the pulldown menu activated by the "Models" tab. Notice that the calcuation using the closed solution is much faster than the finite-different caculation. The parameter set has to be changed separately for each calculation method.
Optimization
If you want to optimize the parameters of the newly selected experiments
- Click on the Optimizer tab (at the bottom of the left pane)
- Change all the DataSet entries under the "Data to Match" heading.
- Click on the Dataset entries to get a selection of data sets to choose from.
- Click on the Curve entries to get a selection of data columns to choose from.
- Hit the "Run" button.
We welcome comments and feedback for this model. Please use the button below to send comments:
- Goresky CA, Bach GC, Nadeau BE. On the uptake of materials by the intact liver. The concentrative transport of rubidium-86. J Clin Invest 52:975-990, 1973.
- Goresky CA, Bach GC, Nadeau BE. On the uptake of materials by the intact liver. The transport and net removal of galactose. J Clin Invest 52:991-1009, 1973.
- Goresky CA, Daly DS, Mishkin S, Arias IM. Uptake of labeled palmitate by the intact liver: role of intracellular binding sites. Am J Physiol 234:E542-53, 1978.
Author: Andreas J. Schwab (andreas.schwab@mcgill.ca)
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

