Linear Flow Coupling. Model I. Const Capillary Transit Time (TT) and Varying Large Vessel TT. Model II. Constant Large Vessel TT and Varying Capillary TT. Model III: Linear relation between capillary and large-vessel TT.
Model number: 0159
Linear Flow Coupling
To model the movement of substances through an organ, the movement along a single path of blood flow through the microcirculation is considered first.
Each flow path consists of two portions: a capillary-bed portion, where exchange with extravascular space occurs, and a noncapillary-bed portion consisting of arteries, arterioles, venules, and veins where all indicators are confined to the vasculature. In linear systems, the order of serial connection is irrelevant, such that arteries, arterioles, venules, and veins (the large vessels) can be lumped together and described by a single distribution of transit times.
The tracer concentration at the outflow of the organ is the flow-weighted average of the outflow concentrations from all the single flow paths. Flow paths of given total transit times, τ, vary in capillary transit time, τc, according to the conditional probability distribution, P(τc|τ).
The following special cases can be considered:
- uniform capillary transit time independent of τ, with variable large-vessel transit time
- variable capillary transit time τc = τ – t0 with uniform large-vessel transit time, t0.
- variable capillary and variable large-vessel transit times
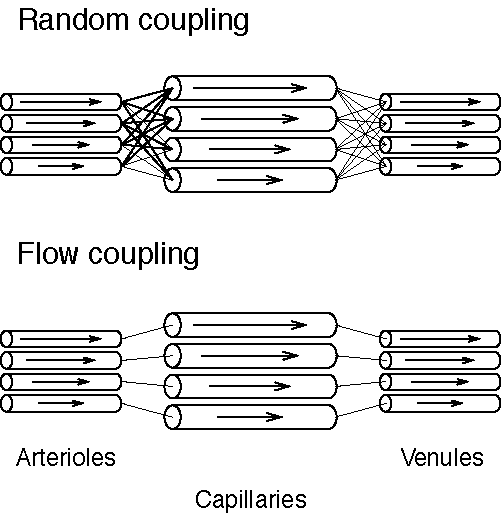
- random coupling: capillary and non-capillary transit times are stochastically independent from each other
- flow coupling: for each total flow-path transit time τ, there is a single capillary transit time τc
- linear relation between capillary and large-vessel transit times
- non-linear relation between capillary and large-vessel transit times
Theory
Model I: Constant Capillary Transit Time and Varying Large Vessel Transit Times
For the intravascular tracer (albumin), the output concentration is
Cref(t) = (q/Fr(tc) = (q/F) r(τl)
where r(τl) is the distribution of large vessel transit times, τc is the common capillary transit time, q is the total amount of tracer injected, and F is total coronary flow.
For the exchanging diffusible tracer: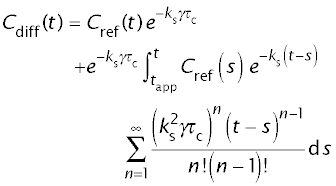
Model parameters to be fitted are:
- ks, the permeability-surface product for efflux from and influx into the capillary per volume of accessible extravascular space
- Φ = 1/(γ τc), the flow per volume of accessible extravascular space
tapp is the time when the label first appears at the exit of the organ, and γ is the ratio of the volume of accessible extravascular space to that of the vascular space.
Model II. Constant Large Vessel Transit Time and Varying Capillary Transit Times
This case is the same as the barrier-limited case for the liver.
For the reference tracer Cref(t) = (q/F) n(t – τl) = (q/F) n(τc) where n(τc) is the distribution of capillary transit times, and τl is the common large vessel transit time.
For the diffusible tracer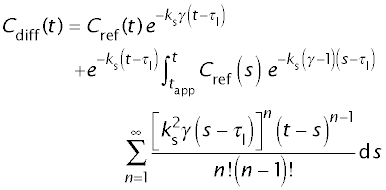
Model parameters to be fitted are:
- ks, the permeability-surface product for efflux from and influx into the capillary per volume of accessible extravascular space
- γ, the ratio of the volume of accessible extravascular space to that of the vascular space
- τl, the large vessel transit time
τcm = tapp – τl is the minimal capillary transit time, where tapp is the appearance time of the tracer.
Model III: Linear relation between capillary and large-vessel transit time
The capillary transit time of a flow path with transit time t is
τc(t) = τcm + b(t – τcm – τlm )
and the large-vessel transit time of a flow path with transit time t is
τl(t) = τlm + (1 – b)(t – τcm – τlm)
where τcm and τlm are the minimal capillaray and large vessel transit times, respectively, with τcm + τlm = tapp.
For the reference tracer Cref(t) = (q/F) w(t – τl – τlm) where w(τc) is the distribution of capillary transit times.
For the diffusible tracer
Substitution of τc and τl yields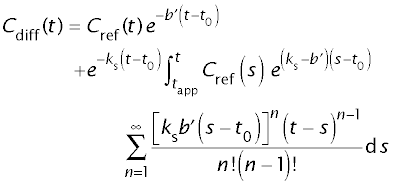
where t0 = (1-1/b)τcm + τlm = tapp – a´/b´ and b´ = ks γ b.
Model parameters to be fitted are:
- ks, the permeability-surface product for efflux from and influx into the capillary per volume of accessible extravascular space
- a´ = ks γ τcm
- b´ = ks γ b.
Relation between the models
Model III can be formulated as a reparametrization of Model II, such that b´/ks in Model III is equivalent to γ in Model II, and t0 in Model III is equivalent to τl. Model II is a realization of the data only if the fitted value for τl> 0. If τl < 0, only Model III can be applied to the data. In the latter case, the heterogeneity of the capillariy transit times is bounded by
b < (tappb´)/a´
and the extravascular/vascular volume ratio is bounded by
γ > a´/(tapp ks)
Experimental design
The following tracers were injected into the coronary artery of an anesthetized dog:
- 125I-albumin (as an indicator for the vascular space)
- [U-14C]sucrose (as an indicator for the combined vascular and interstitial spaces)
Samples obtained at the coronary sinus were analyzed for beta and gamma radioactivity.
With each dog, a control injection (run 1) was performed, followed by a second injection (run 2) performed in a vasodilated state. In Experiment 1, vasodilation was achieved by intracoronary infusion of flavaspidic acid. In Experiment 6, vasodilatation was achieved by intracoronary infusion of 2-bromopalmitate.
Download JSim model project file
| Run Model: | Help running a JSim model. |
(Java model applet may take 10-20 seconds to load.)
There are twelve parameter sets in this model project file, for two experiments, two runs per experiments, and three model evaluation for each run.
To change the parameter set:
- Select Load project parameter set from the ParSet pulldown menu
- Choose the desired parameter set. This will automatically change the paremters as well as the data for the reference curve.
- Click on "Run" to use the new parameter set.
To plot the results of the simulations, the appropriate data sets and models have to be chosen in the plot pane.
- Select a cuve from the pulldown menu labeled "Curve"
- Select a data set or a model from the pulldown menu labeled "Data"
If you want to optimize the parameters of the newly selected experiments:
- Click on the Optimizer tab (at the bottom of the left pane)
- Change all the DataSet entries under the "Data to Match" heading.
- Click on the Dataset entries to get a selection of data sets to choose from.
- Click on the Curve entries to get a selection of data columns to choose from.
- Hit the "Run" button.
Model Feedback
We welcome comments and feedback for this model. Please use the button below to send comments:
References
CP Rose and CA Goresky. Vasomotor control of capillary transit time heterogeneity in the canine coronary circulation. Circ. Res. 39:541-554, 1976
Author: Andreas J. Schwab (andreas.schwab@mcgill.ca)
Related Models
Back to Goresky Modeling of transport and metabolism tutorial
- Flow limited model
- Simple elimination with flow-limited distribution
- Barrier limited model
- Linear flow coupling
- Two-barrier model
- Red cell model
Key Terms
indicator dilution, flow-limited, liver, transport, vascular volume, organ, disse space, Goresky transport tutorial
Acknowledgements
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send an email with the citation and, if possible, a PDF file of the paper to:
Or send a copy to:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

