Simple elimination with flow-limited distribution
Simple elimination with flow-limited distribution
A tracer that is rapidly equilibrated between the vascular an extravascular space and is eliminated from this space will exhibit flow-limited behavior with elimination.
Sulfobromophthalein
Sulfobromophthalein (BSP) is an organic dye used a for liver diagnostic. Its chemical structure is illustrated by the following formula:
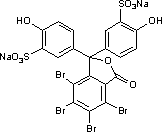
BSP binds tightly but reversibly to albumin and therefore occupies the same space. Together with albumin, it distributes rapidly between the vascular space and the interstitial space. It is eliminated from the interstitial space by uptake into hepatocytes.
Experimental design
The following tracers were injected into the portal vein of an anesthetized dog:
- Erythrocytes (Ery) labeled with 51Cr
- Evans Blue (Alb) (a tracer for albumin)
- Sulfobromophthalein (BSP) labeled with 32S
Samples from a hepatic vein were analyzed for radioactivity and spectrophotometrically for Evans blue.
BSP was injected in various doses. In this tutorial only two experiments are demonstrated with doses of 0.06 mg and 6.61 mg.
Calculations
Albumin exchanges rapidly between the vascular and the interstitial space (the space of Disse). Therefore, it is shows delayed-wave transport along the sinusoids that follows the equation derived in the [[Transport Physiology:Whole organ models:Goresky:Flow_limited_model:model_index|chapter on "flow-limited distribution"]]:
| (1) | Cref(t) = | 1
1 + γref |
Cv( | t − t0
1 + γref |
+ t0) |
The BSP curve is calculated using the equation
| (2) | C(t) = Cappref(t) e−k(t − t0) |
where k is the elimination coefficient. Cappref is the outflow profile of a hypothetical reference indicator that occupies the same space as BSP but is not eliminated and whose behavior is described by an equation similar to that for albumin:
| (3) | Cref(t) = | 1
1 + γ |
Cv( | t − t0
1 + γ |
+ t0) |
|
There are seven parameter sets in this applet:
- BSP1: Sulfobromophthalein experiment with low injected amount
- BSP2: Sulfobromophthalein experiment with high injected amount
- EtOH_1: Ethanol experiment with no ethanol infused
- EtOH_2: Ethanol experiment with low ethanol concentration
- EtOH_3: Ethanol experiment with high ethanol concentration
- PrOH_1: Propanol experiment with no ethanol infused
- PrOH_2: Propanol experiment with high ethanol concentration
To change the parameter set:
- Select Load project parameter set from the ParSet pulldown menu
- Choose the desired parameter set. This will automatically change the paremters as well as the data for the reference curve.
- Click on "Run" to use the new parameter set.
- Select the appropriate plot page from the Plot Pages pulldown menu.
If you want to optimize the parameters of the newly selected experiments
- Click on the Optimizer tab (at the bottom of the left pane)
- Change all the DataSet entries under the "Data to Match" heading.
- Click on the Dataset entries to get a selection of data sets to choose from.
- Click on the Curve entries to get a selection of data columns to choose from.
- Hit the "Run" button.
Monohydric alcohols (ethanol and propanol)
Experimental design
The following tracers were injected into the portal vein of an anesthetized dog:
- Erythrocytes (Ery) labeled with 51Cr
- Water (H2O) labeled with 3H
- Ethanol or propanol labeled with 14C
Samples from a hepatic vein were analyzed for radioactivity.
In some experiments, diluted ethanol was infused peripherally for 30 min before and during the multiple-indicator dilution experimental runs at 1 ml/kg body wt over 30 min.
Calculations
Water exchanges rapidly between the vascular and the extravascular space, the latter comprised of the extracellular space and the interior of the parenchymal cells. Therefore, it is shows delayed-wave transport along the sinusoids, and Equ. 1 applies, where γe is defined as the ratio of the extravascular to the vascular space, given by:
| (4) | γe = | 1 + β + γ + θ
1 + β |
The parameters β, γ, and θ represent the ratio of the accessible space in erythrocytes, interstitium, and parenchymal cells, respectively, to that available in vascular plasma.
Ethanol and propanol also exchange rapidly between the vascular and the extravascular space. Additionally, due to their lipid solubility, the apparent parenchymal space, θ´, is larger than the corresponding water space, θ. The hypothetical reference indicator (dashed line in the JSim plots) is described by Eq. 3, where γref is defined by:
| (5) | γref = | 1 + β + γ + θ′
1 + β |
The observed outflow profile for ethanol or propanol is then given by Equ. 2 with
| (6) | k = | Vmax&theta
(1 + β + γ + θ′)(û + Km) |
where Vmax is the maximal elimination velocity per unit parenchymal space, Km is the Michaelis constant, and û is the logarithmic average of the ethanol concentration in plasma. The latter is defined as:
| (7) | û = | Cin − Cout
ln Cin − ln Cout |
where Cin and Cout are the hepatic inflow (flow-averaged arterial and portal venous) and outflow (hepatic venous) ethanol concentrations, respectively.
In the absence of ethanol infusion, saturable binding to alcohol dehydrogenase results in an additional "enzymic" distribution space with an apparent volume of θEt/Km, where θEt is the total concentration of binding sites and Km is their affinity to ethanol. This increases the value of γref to:
| (8) | γref = | 1 + β + γ + θ′ + θEt/Km
1 + β |
and, accordingly,
| (9) | k = | Vmax&theta
(1 + β + γ + θ′ + θEt/Km )(û + Km) |
We welcome comments and feedback for this model. Please use the button below to send comments:
- Goresky, CA. Initial distribution and rate of uptake of sulfobromophthalein in the liver. Am J Physiol. 207: 13-26, 1964.
- Goresky CA, Gordon ER, Bach GG. Uptake of monohydric alcohols by liver: demonstration of a shared enzymic space. Am J Physiol. 244:G198-214,1983.
- Goresky CA, Bach GG, Rose CP. Effects of saturating metabolic uptake on space profiles and tracer kinetics. Am J Physiol. 244:G215-32, 1983.
Author: Andreas J. Schwab (andreas.schwab@mcgill.ca)
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

