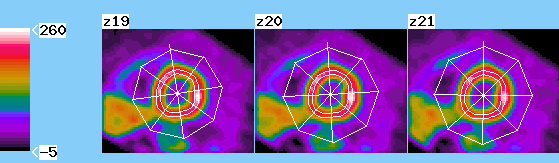
I4 System Overview
I4 is an open software system for biomedical image processing that explicitly support 4 dimensions (3 spatial dimensions and time) as is required by modern imaging technologies such as Positron-Emission tomography (PET) and Magnetic Resonance Imaging (MRI). It is designed as an open system, so that researchers may easily add modules specific to their own research that will work cooperatively with the base functionality of the system.
Biomedical imaging functionality is often frustrated by monolithic commercial systems that are missing a key component needed for specialized research. I4 allows a simple path toward integration of such individualized components.
Key I4 system features include:
- Support for XYZT Pixel images, ROIs, cardiac "bullseye" images, and time activity curves
- A host of well documented, simple utilities for routine image processing tasks such as ROI sampling, decay correction, tomograph correction, image arithmetic, etc.
- A efficient and powerful scripting language for image processing functions that allows researchers to add additional utilities with minimal effort
- A pipelining methodology for combining base system and user-written utilities
- A scripting methodology for the same thing, but performing more quickly
- Support for set-theoretic ROI operations, allowing complex sampling strategies
- Intelligent, uncluttered, and open file specifications with documented access libraries allowing researchers to write additional programs without "reinventing the wheel"
- An interactive image viewing query program supporting image overlay, custom color scales and point-and-click 3-D ROI editing
- AI utilities for automatic definition of ROIs in image sets (journal article in preparation)
- Interfaces to modelling programs for functional imaging
- A commitment to open software systems
- Fully available source code
The I4 system is now available for download via the internet.
System Design and Philosophy
I4 grew out of our need for an open system capable of dealing with 3D and 4D image files. A number of commercial packages deal with such images, but they are uniformly expensive and closed in their design, making them problematic for ongoing research projects. Several open-design public-domain packages deal with 2D image manipulation, most admirably Jef Poskanzer's PBM. Unfortunately, dealing with 3D and 4D image sets within the confines of a 2D package becomes a more and more arduous collation problem, rather like doing numeric integration on a 4 function pocket calculator.
I4 attempts to be "the PBM of biomedical imaging". The PBM (now PBMPLUS) package, originally written by Jef Poskanzer, is a set of 2-D imaging utilities whose great utility is due to
- simple, intelligent, portable file specifications
- a base set of simple, portable tools
- a reasonable framework for adding new tools
- a pipeline design that facilitates combining old and new tools
PBM does not do all things for all people. It provides a base functionality and a framework for creating custom tools that can be combined easily with the base set. Users can do original work without having to reinvent the wheel.
I4 attempts to extend the great design properties of PBM to the 4-D imaging requirements of such technologies as Positron-Emission tomography (PET) and Magnetic Resonance Imaging (MRI). Programs are written to be highly portable. File specifications are independent of underlying hardware or operating system. Simple command line tools exist for a base set of processing functions which may be combined in pipeline or scripts to perform more complex tasks. Standards are defined so that researchers may easily add custom tools that work coorperatively with the base set.
Many commercial imaging systems are designed as large monoliths will many disparate functions packed into a labyrinthian, point-and-click front end. I4 instead does most of its image processing as command line scripts. Interactive functions such as image display and manual ROI editing take place in a smallish viewing program (i4view). This is not because point-and-click is "bad", rather
- much image processing requires (sometimes complex) protocols be established for routine processing of large datasets, scripting is more efficient that point-and-click in such environments
- great programmer effort is required to make point-and-click interfaces for functions that do fine on the command line, effort that would be better spend improving system functionality
- point-and-click applications cannot be pipelined or easily combined
- monolithic programs usually become opaque and difficult to maintain
- monolithic programs discourage cooperative research by preventing integration of user-added utilities
Like PBM and unlike commercial imaging systems, I4 does not spend great effort keeping a large database of labelling information pertaining to an image file. Such information is best tracked via database technology external to the image files themselves. By eliminating database information from the image files, I4 processing programs become simpler to read and easier to write.
Further Information
The base I4 system was designed and written by Erik Butterworth.
Copyright ©: 1998-2007 National Simulation Resource, University of Washington, Seattle, WA 98195-7962.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

