In a single compartment with flow, substrates Xa (Xanthine) and Ua (Uric acid convert to each other.
Description
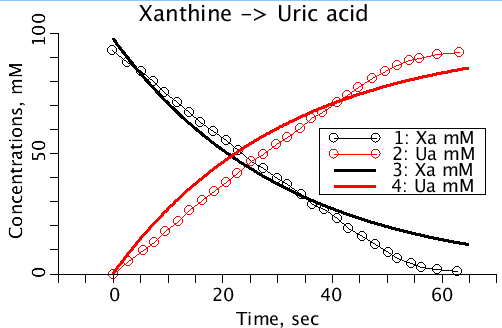
This is a one compartment model with flow and a reaction. In the absence of flow or other reactions (with no back reaction) the situration modeled is a "Progres" experiment as Xa converts to Ua. The file contains a data set on the conversion of Xanthine to Uric Acid taken from Escribano et al 1988. The program includes a back reaction, Uric Acid to Xanthine. Optimizing on Flow, the initial concentration of Xanthine, and the two reaction rates sets both flow and the backwards reaction rate to zero. The plots of the data and the model curves show that this model does not describe the measurements. The reaction rate appears to be too fast initially and too slow later. The misfitting relegates the first order reaction model used here to the dustbin. See models 320, 321, 322, 323, 324.
Equations
Ordinary Differential Equations
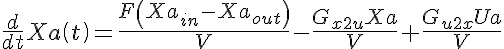
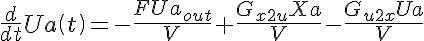
Initial Conditions
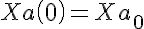
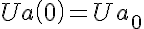
The equations for this model may be viewed by running the JSim model applet and clicking on the Source tab at the bottom left of JSim's Run Time graphical user interface. The equations are written in JSim's Mathematical Modeling Language (MML). See the Introduction to MML and the MML Reference Manual. Additional documentation for MML can be found by using the search option at the Physiome home page.
- Download JSim model MML code (text):
- Download translated SBML version of model (if available):
We welcome comments and feedback for this model. Please use the button below to send comments:
Bassingthwaighte James B., Chinn Tamara Meiko, Re-examining Michaelis-Menten enzyme kinetics for xanthine oxidase, Adv Physiol Educ 37: 37-48, 2013 Escribano, J., Garcia-Canovas, F., and Garcia-Carmona,F. A kinetic study of hypoxanthine oxidation by milk xanthine oxidase. Biochem. J. 254: 829-833, 1988. Michaelis L and Menten ML. Die Kinetik der Invertinwirkung. Biochem Z 49: 333-369, 1913.
Please cite https://www.imagwiki.nibib.nih.gov/physiome in any publication for which this software is used and send one reprint to the address given below:
The National Simulation Resource, Director J. B. Bassingthwaighte, Department of Bioengineering, University of Washington, Seattle WA 98195-5061.
Model development and archiving support at https://www.imagwiki.nibib.nih.gov/physiome provided by the following grants: NIH U01HL122199 Analyzing the Cardiac Power Grid, 09/15/2015 - 05/31/2020, NIH/NIBIB BE08407 Software Integration, JSim and SBW 6/1/09-5/31/13; NIH/NHLBI T15 HL88516-01 Modeling for Heart, Lung and Blood: From Cell to Organ, 4/1/07-3/31/11; NSF BES-0506477 Adaptive Multi-Scale Model Simulation, 8/15/05-7/31/08; NIH/NHLBI R01 HL073598 Core 3: 3D Imaging and Computer Modeling of the Respiratory Tract, 9/1/04-8/31/09; as well as prior support from NIH/NCRR P41 RR01243 Simulation Resource in Circulatory Mass Transport and Exchange, 12/1/1980-11/30/01 and NIH/NIBIB R01 EB001973 JSim: A Simulation Analysis Platform, 3/1/02-2/28/07.

